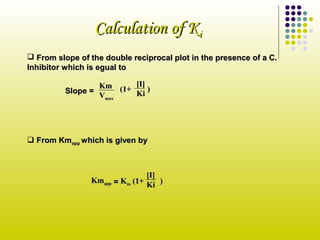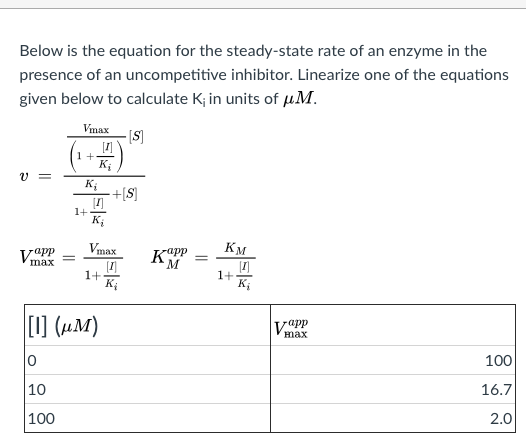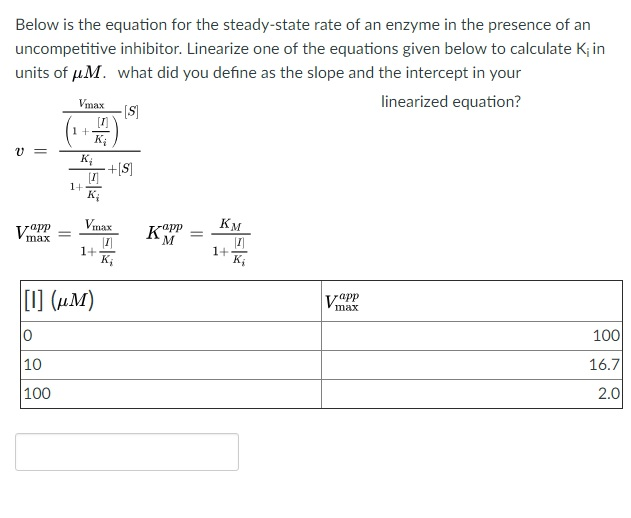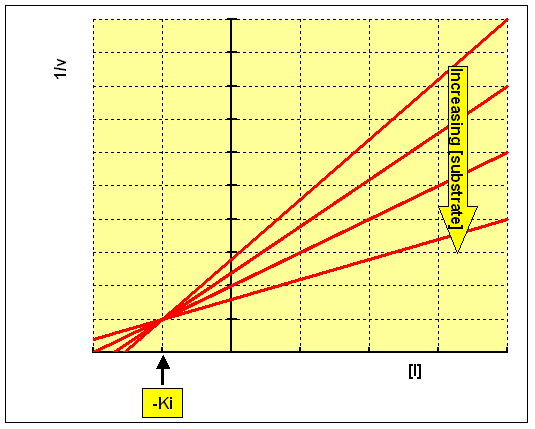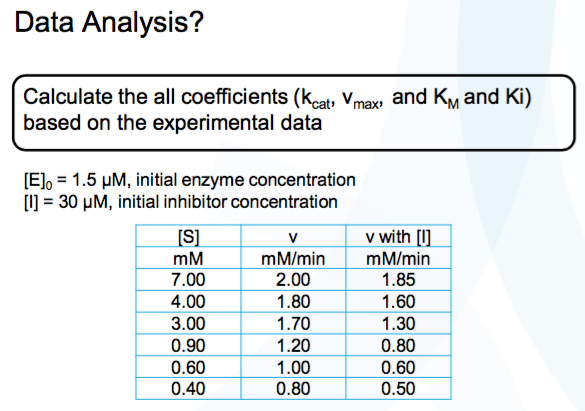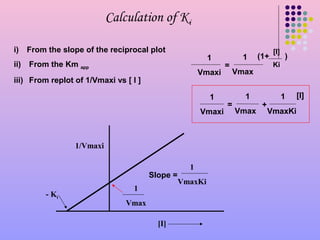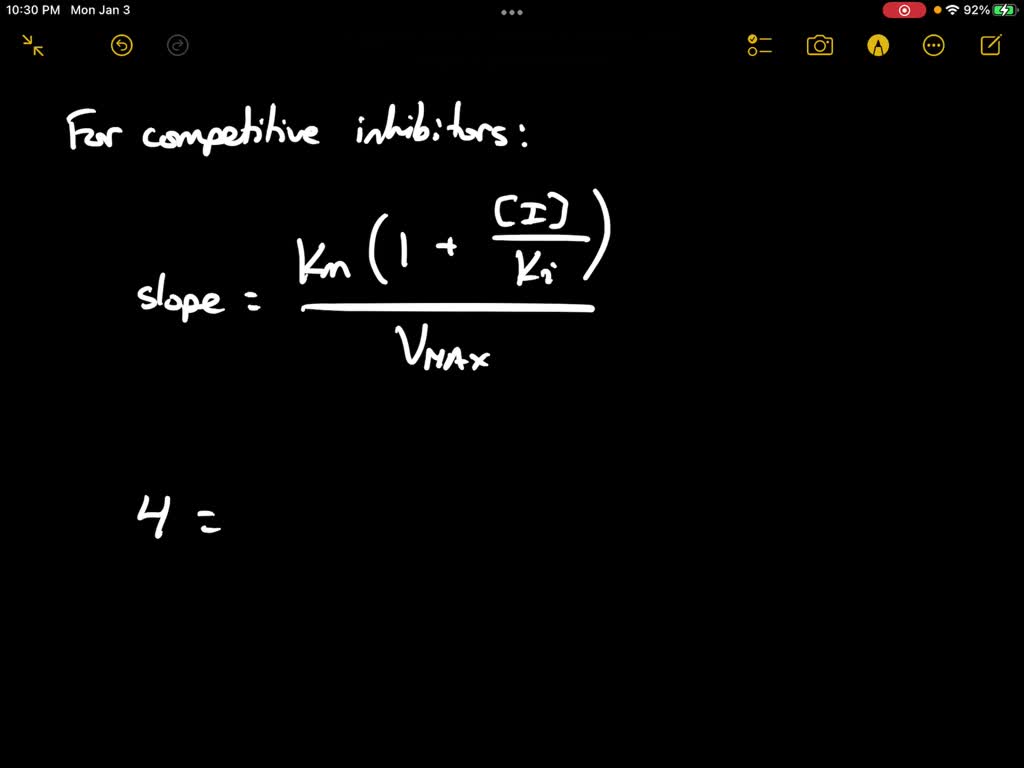
SOLVED: Calculate the Ki for a competitive inhibitor whose concentration = 200 mg/mL, Km = 0.80, vmax = 0.20, slope = 4. Please show work.

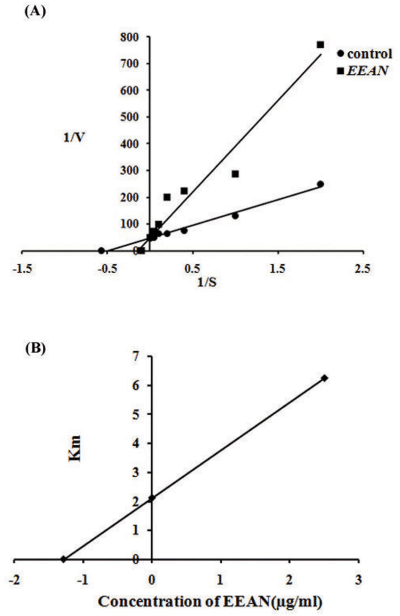
Lineweaver-Burk secondary plot for Ki calculation activity. Compound 3b... | Download Scientific Diagram
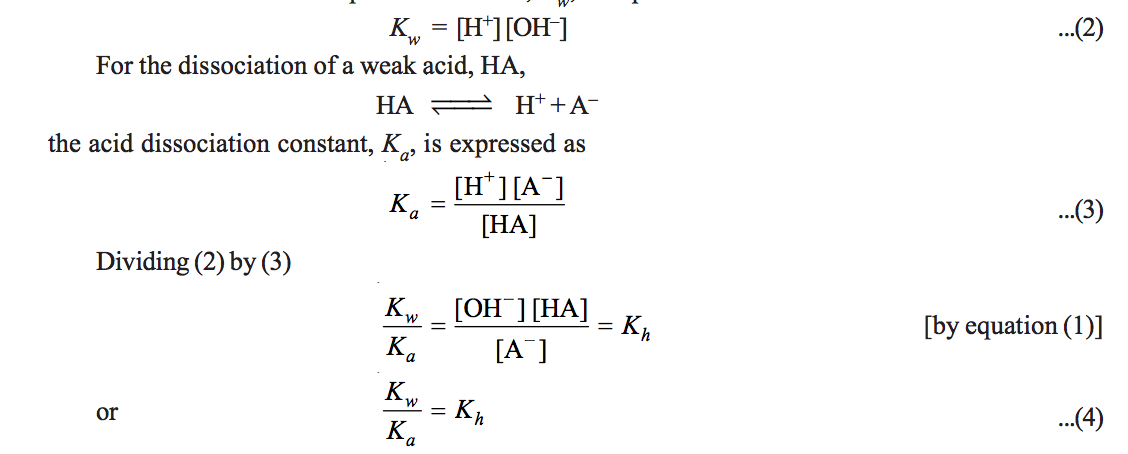
Calculation of Hydrolysis Constant, Degree of Hydrolysis and pH of Salt Solution - Chemistry, Class 11, Ionic Equilibrium

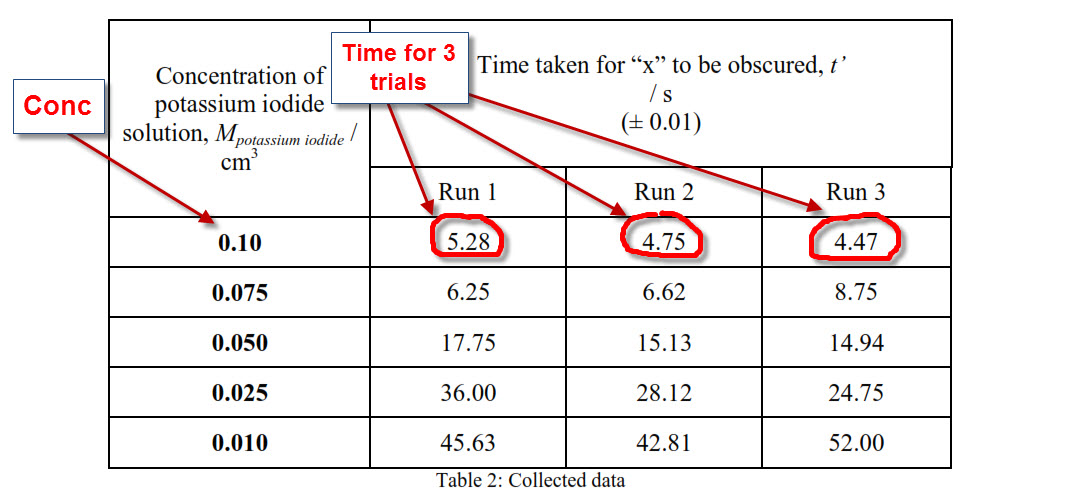
Calculate K(c ) for the reaction KI+I(2) hArr KI(3). Given that initial weight of KI is 1.326 g weight of KI(3) is 0.105 g and number of moles of free I(2) is

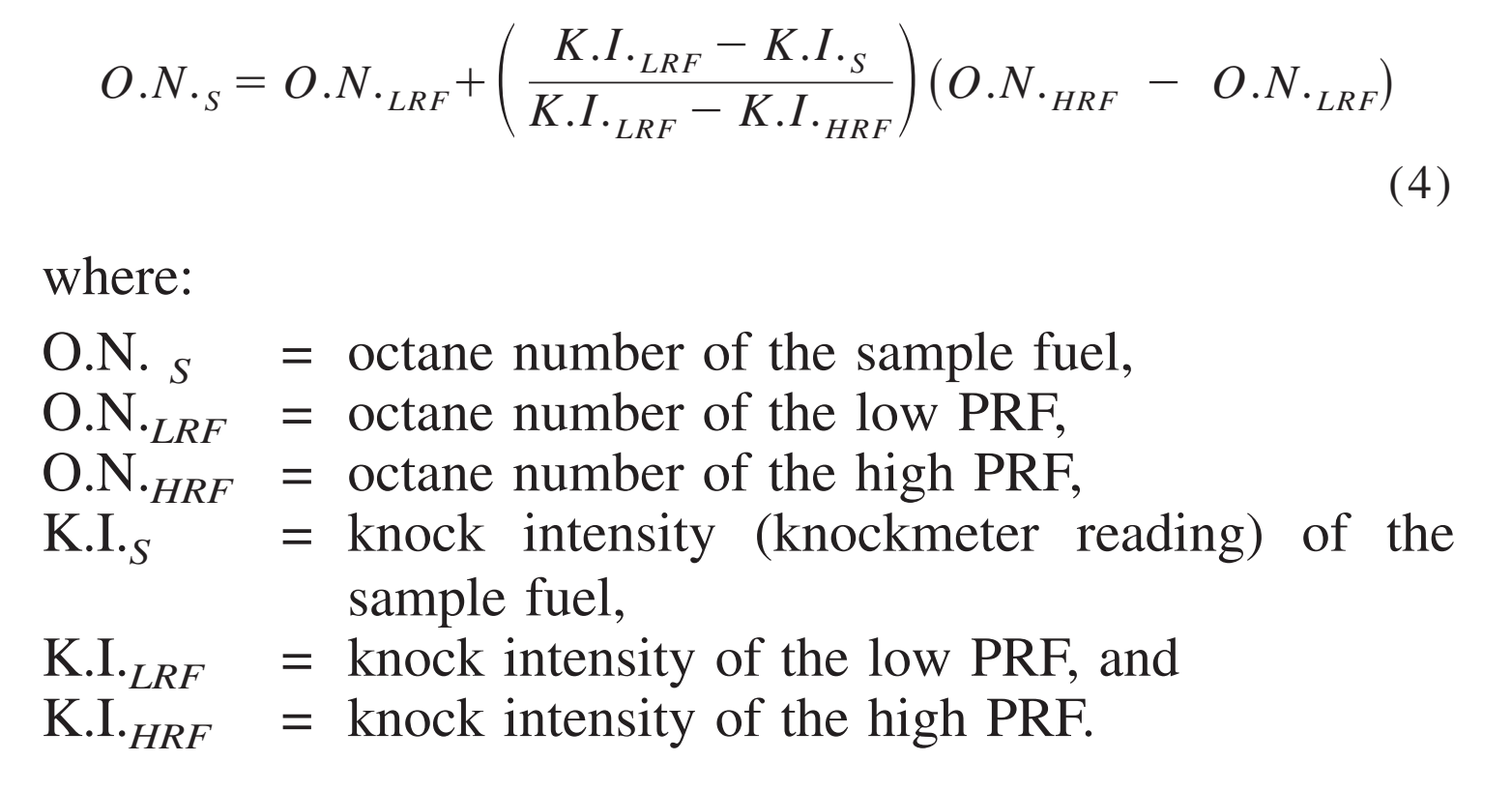
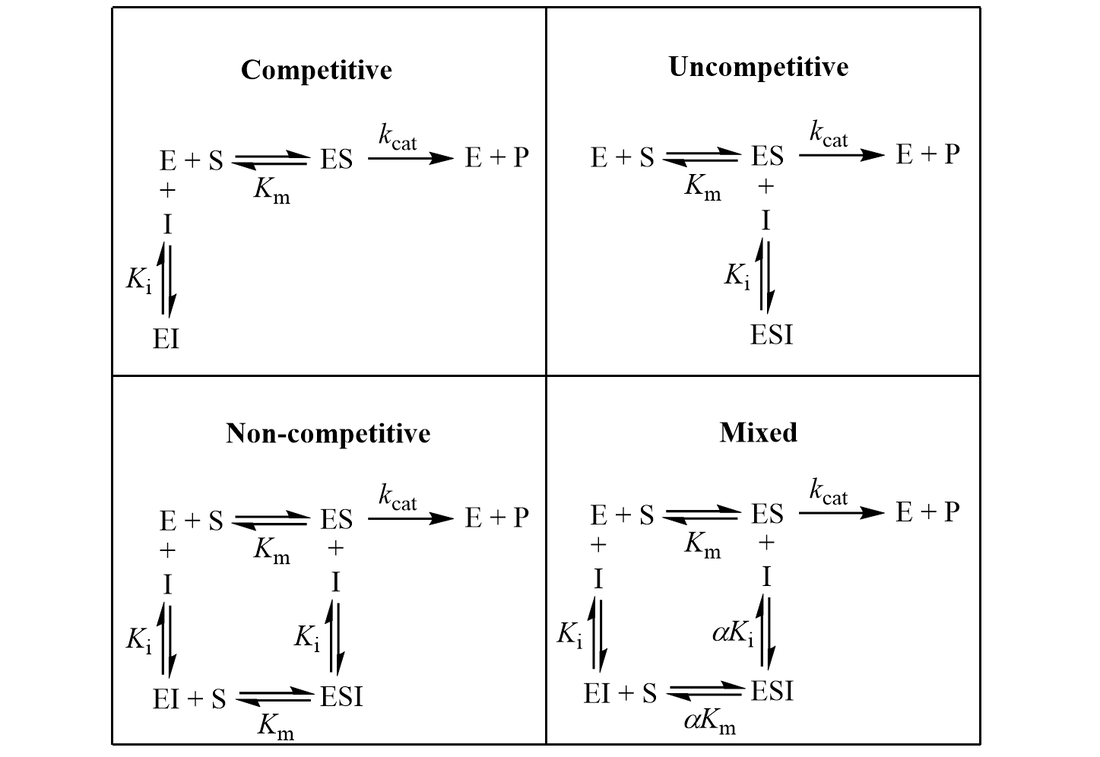
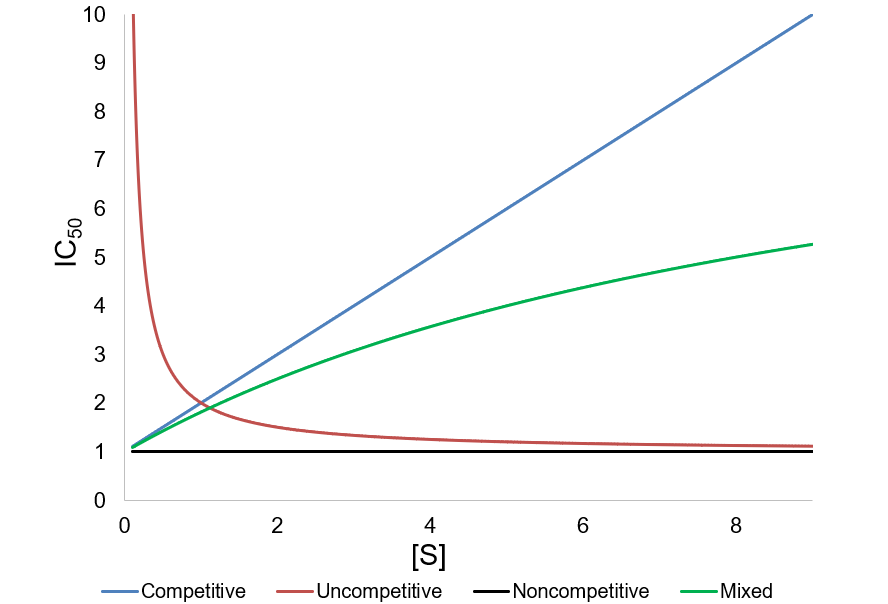
A METHOD FOR IDENTIFICATION OF INHIBITION MECHANISM AND ESTIMATION OF KI IN IN VITRO ENZYME INHIBITION STUDY | Drug Metabolism & Disposition